Application of Visualization and Haptic Feedback to Enhance Molecular Docking
Principal Investigator:
Dr. David R. Bevan,
Department of Biochemistry
Co-Principal Investigators:
Dr. Layne T. Watson,
Department of Computer Science
Dr. Ronald D. Kriz,
Department of Engineering Science and Mechanics, and
Director of the University and Visualization Group
("VT-CAVE")
Ph.D. Graduate Student, Biomechanics, Mr. Sanjiv Parikh
Project Description
Molecular docking, the process by which two or more molecules recognize and interact
with one another, is a crucial event in biochemistry. For example, it is central to
many fundamental regulatory processes in cells, and it is the basis for practical
applications such as drug design and protein engineering. Major challenges in this
field are (1) predicting how molecules will dock with one another and (2) identifying
the factors that determine the specificity of interaction.
Several approaches to molecular docking have been developed, all of which rely on
the number-crunching capabiliities of computers to generate and evaluate large numbers
of possible binding orientations. However, they do not take advantage of the
graphical capabilities offered by computers. Moreover, existing methods purportedly
enhance objectivity by limiting human intervention, but in so doing they severely
limit the scientist's ability to analyze the solutions that are provided and exploit
their chemical intuition, expertise, and experience in generating a sound and
reliable solution.
To make major advances in molecular docking, it is necessary to
develop novel methods to facilitate the docking techniques by enhancing our ability
to understand and analyze docking interactions and to develop hypotheses about which
molecules are most likely to interact favorably. These factors can be incorporated
into molecular docking by including visualization and haptic (sense of touch)
feedback in the process. With this funding from ASPIRES, we will combine visualization
and haptic feedback to study molecular docking on workstations, and Immersadesk,
and a completely immersive environment, the CAVE. As a result, we will have at our
disposal a state-of-the-art research tool that is not available at most other
institutions, thereby increasing our competitive advantage for external funding.
Specific Objectives
- To implement haptic feedback for interactive molecular docking on graphics
workstations and in the CAVE.
- To evaluate the effectiveness of interactive docking by comparing the results of
docking simulations with experimental data.
- To perform docking simulations, the results of which will support current research
as well as provide preliminary data for grant applications.
- To evaluate the ease of use of interactive molecular docking.
Progress (March 17, 2001):
PHANToM <->
VRPN <->
DIVERSE
link between Windows PCs and Linux workstations, shown in Figure 2, has been successfully
demonstrated. The next step is to include the CAVE and IDesk with DIVERSE. Expected
completion date May 2001.
Progress (August 28, 2001):
Final report (NISTreport.doc) and associated code can be downloaded from
NIST_phaseII
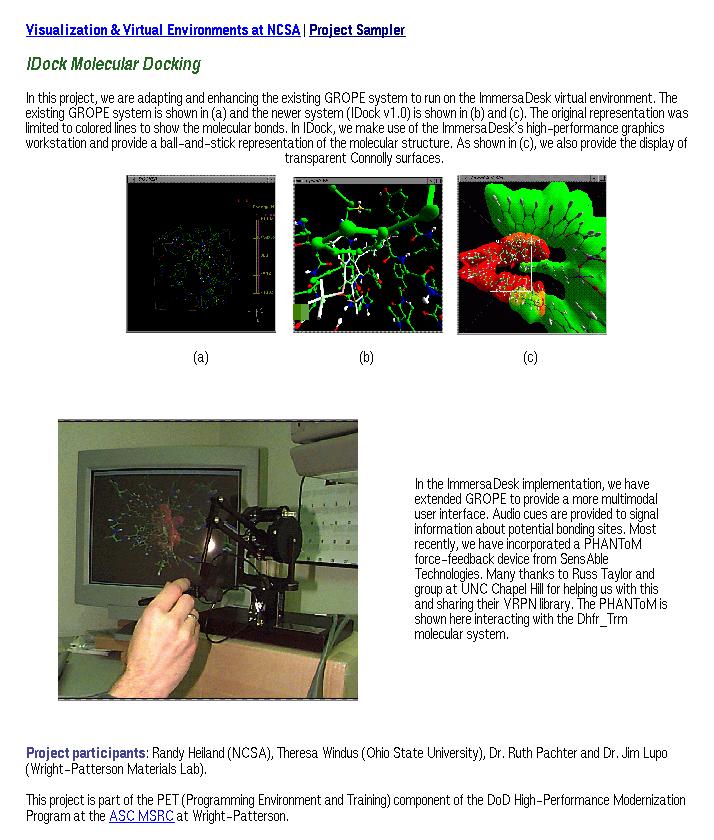
Figure 1. NCSA IDock Project, http://www.ncsa.uiuc.edu/Vis/Projects/Docker
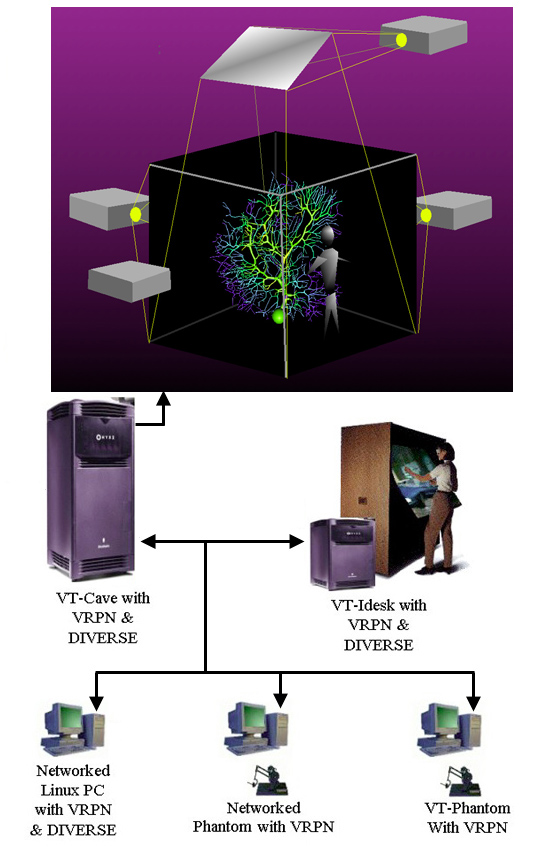
Figure 2. IDock Device Independent Virtual Environemnt.
Select this link for large image of poster
Figure 3. Poster presentation at the Virginia Tech Bioinformatics Conference,
March, 19, 2001.
Please contact David Bevan at
drbevan@vt.edu
or Ron Kriz at:
rkriz@vt.edu for more information
Last Revision March 15, 2001
http://www.sv.vt.edu/future/cave/resprj/idock/idock.html

